Clinicians and medical researchers believe that healthcare systems would enhance their performance if integrative diagnostic approaches were implemented. Although these approaches are advantageous, they are time-consuming, expensive and require complex interpretation, making it hard to be implemented in clinical laboratories. In the last decade, high-throughput technologies such as NGS, microarrays, RNAseq and MALDI-ToF have been evolving and demonstrating in research studies to have an enormous potential to be applied as integrative approaches in personalised medicine and clinical diagnostics.
With these technologies fully implemented in clinical laboratories, it would be possible to extract a large amount of information about the genome, transcriptome, proteome, metabolome and phenome of patients. The information available from these “omics” data is rich enough to allow screening and early detection of multiple diseases as well as the detection of therapeutic targets in drug discovery. High-throughput technologies have been improving the rate by which they generate data from biological samples in such a way that become acceptable for clinical application and personalised medicine.
For example, MALDI-ToF is an ultra-fast and affordable high-throughput technology that processes multiple samples in the scale of minutes, rendering mass spectra that contain proteomic and/or metabolomic information. NGS and RNAseq, on the other hand, are two high-throughput technologies that give robust information about the sequence of the genes and its expression level, which would make possible to detect mutations and deregulation’s associated to genetic diseases. In some clinics, MALDI-ToF is already been used for the detection of bacterial strains, whereas NGS, microarrays and RNAseq for detection of some genetic diseases. Most of these high-throughput technologies have a high cost in the beginning with the acquisition and implementation in laboratories. However, the cost of operations is now quite affordable, particularly in the case of MALDI-ToF, which would pay off the investment and optimise the analytical service in a long-term scale. Moreover, these high-throughput technologies should be seen as potential tools for screening multiple diseases from one patient’s biological sample, which would further decrease the operational costs per disease if proper analytical systems are implemented.
Bioinformatics as a solution for clinical diagnostics
Integrating a large amount of data coming from high-throughput technologies towards personalised medicine and diagnostics cannot be possible without using computational approaches to sort out the complexity of processing and correlating multiple variables at the “omics” level. Bioinformatics is an interdisciplinary field of biology that is focused on applying computational techniques for the analysis and extracting information from data coming from biomolecules. Usually, it integrates techniques from the fields of informatics, computer science, molecular biology, genomics, proteomics, mathematics, and statistics. Although it started as a field fully dedicated to basic research in evolution and genetics, it has been evolving in parallel with high-throughput techniques resulting in the development of many methods and tools that facilitate the interpretation of “omics” data. In bioinformatics, high-throughput data is processed and analysed systematically from raw data to the results using pipelines of analysis using the full potential of computers. Bioinformatic pipelines usually contain multiple steps for data quality assessment, feature extraction, dimension reduction, biomarker detection and results generation. This set of analysis is fully automated where the user has no interference but can play the role of “curator” to check the validation of the outputs (results).
With the evolution of the computational power, bioinformatics gained the potential to tackle big data and integrate a large amount of data much faster than it is produced, becoming a solution applying high-throughput techniques in clinical diagnostics and personalised medicine. For example, some studies have demonstrated that bioinformatic pipelines developed for the analysis of MALDI-ToF mass spectra can extract diagnostic information from urine, blood and embryo culture media faster than its capacity of being generated. In genomics, several bioinformatic pipelines of analysis for NGS, RNAseq and microarrays have been also developed to extract diagnostic information out of sequencing of virus, pathological bacteria and cancer biopsies.
Moreover, bioinformatics tools for processing “omics” have also been successful in the discovery of novel drug targets for cancer therapy. Bioinformatics can further improve clinical laboratories efficiency and costs by saving time and human resources on the analysis and reporting to clinics and patients. This can be done by developing pipelines of analysis with automated reporting and APIs fully dedicated to giving real-time online access, facilitating the communication between laboratories, clinicians and patients. Besides, patient historical data and metadata should be secure and organised in a structured way (data “warehouses”) such that it can be further pulled systematically to bioinformatics pipelines. This would allow going beyond in integrative analysis of patients by having their data as a function of time allowing a more personalised monitoring of patients diagnostic and allowing better prognostics.
Predictive modelling as a complementary diagnostic tool
Predictive modelling frameworks have been extensively used for describing physiological systems and diseases. These frameworks use mathematical models and algorithms for generating predictions about a phenotype or making reasonable estimates with the available data. The integration of these predictors in bioinformatic pipelines is fundamental to make accurate classifications of patient’s samples into the likelihood of having a particular disease or not. This methodology is useful in clinical laboratories for screening and early detection of genetic or metabolic diseases. Also, this type of approach allows making estimations of inaccessible body chemistry parameters based on others, which otherwise are impossible due to experimental constraints or the available methods are too invasive/expensive.
The development of mathematical models and algorithms that generate robust predictions is a hard task and requires rigorous validation procedures before a predictor is ready to be launched into the market. Not many predictors for diagnostics are available to be used or can be adapted to a given clinical laboratory setting. Thus, model development and optimisation for each lab would be the ideal scenario. Integrating predictive modelling workflows in bioinformatic pipelines also facilitate model development by systematising the process of validation and model selection using the data and metadata.
Several types of models can be used to make diagnostic predictions and the choice depends on the data available, technology and the nature of the problem. Statistical models based on known distributions of biomarkers are very common to be used in the diagnostic of a particular disease. These are easy to implement in bioinformatic pipelines and serve as complementary information for clinicians. Implementation of pattern recognition, machine learning and artificial intelligence (AI) algorithms into bioinformatic pipelines are key to optimise mathematical models towards meeting more accurate predictions. Importantly, the usage of machine learning and AI algorithms are essential for the idea of personalised medicine because they enable the fitting of generic models of disease to each patient scenario and body chemistry. Deterministic models such as the logical and kinetic modelling frameworks can also be used for simulation of physiological scenarios and making robust predictions with clinical applications.
For example, simulation of the tumour micro-environment using a logical network model of the regulation of cell adhesion properties allowed to establish relations between cancer de-regulations and the metastatic potential. This has a huge potential for the future development of bioinformatics tools that allow the prediction of the metastatic potential and suggest the best therapy for each case based on the tumour biopsy. Kinetic models, on the other hand, have the potential to be more precise and generate a continuous range of prediction values. However, their parameter estimation is complex and requires machine learning algorithms to adapt them to a particular physiological system. These types of models are excellent for describing the metabolism and can be very useful in as future tools in personalised medicine.
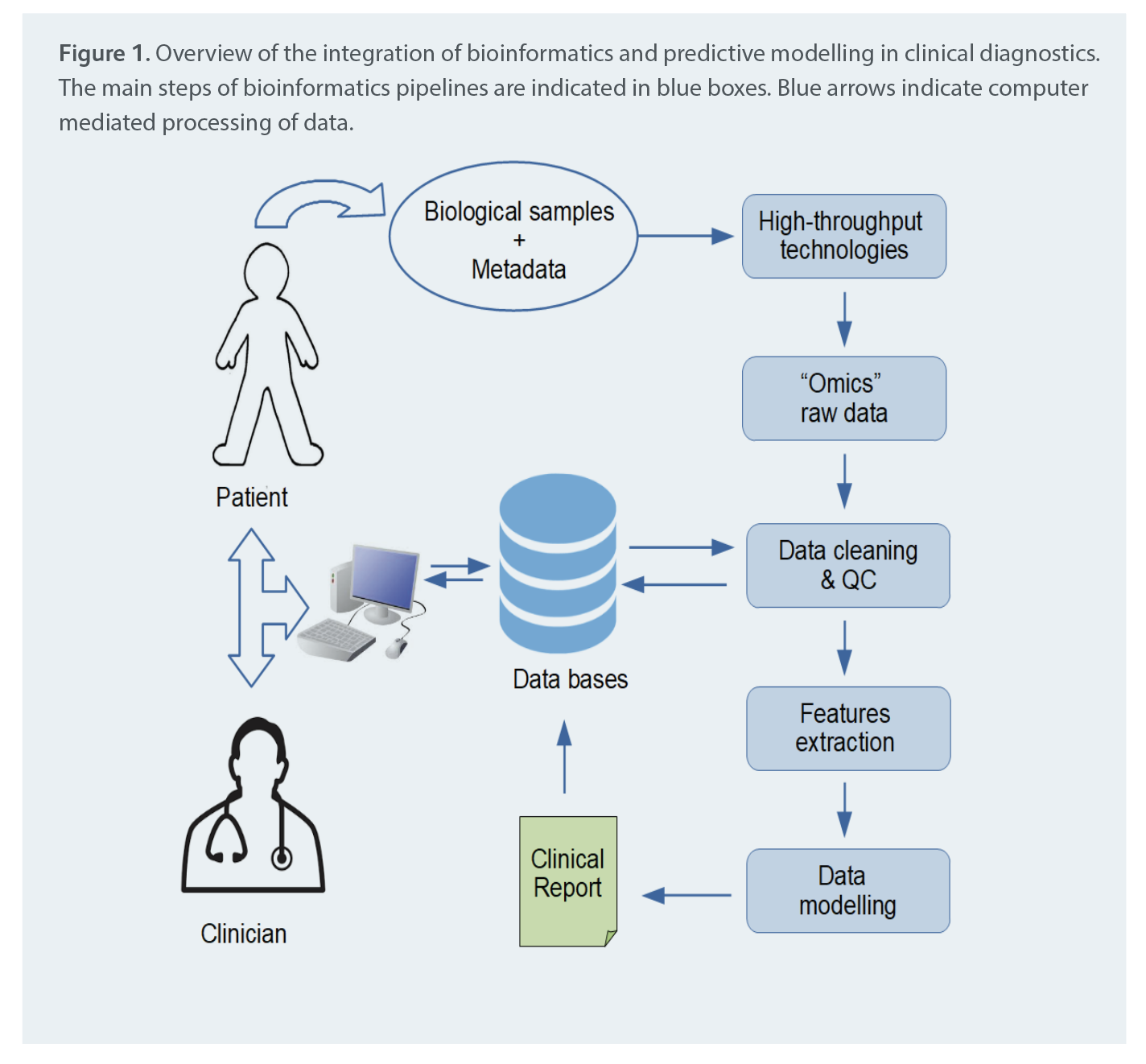
Implementing bioinformatics in clinical laboratories
There is still much to be done for the implementation of the full potential of bioinformatics as a diagnostic tool in clinical laboratories. This requires a joint effort between clinical laboratories, healthcare systems, and software companies to make it happen. As initial steps, clinical laboratories and healthcare systems should start to invest in the following:
- Acquisition of “omics” high-throughput laboratory equipment (NGS, RNAseq and MALDI-ToF).
- Acquisition of computational resources (high-performance computers and servers).
- Hire bioinformaticians or specialised outsourcing companies.
- Implementation bioinformatic tools specifically designed for each laboratory reality.
- Implementation of predictive models in software applications for clinical diagnostics.
- Implementation of web platforms for connecting laboratories, patients and clinicians.
Nevertheless, health organisations should also make an effort to recognise, legislate and validate most bioinformatics and predictive modelling diagnostic tools. Although, this would be a big investment in time, money and resources. However, in the future, it would pay off in terms of the quality of the diagnostic services offered to the populations, making it evolve towards personalised medicine that is affordable for most people.
References available on request.


